

STANDING



OF TIME
Marking 100 years of the JAX Summer Student Program
THIS ISSUE CELEBRATES
a milestone in JAX’s history: the centennial of our Summer Student Program. In 1924, five years before the Laboratory was officially established in 1929, JAX founder Clarence Cook Little began bringing small groups of students to Mount Desert Island, Maine, to conduct summer research projects.
Over the last 100 years, what became the JAX Summer Student Program (SSP) has brought several thousand high school and college students to Bar Harbor and, for the past decade, Farmington, Connecticut, to conduct original research alongside postdoctoral and faculty mentors. The experience has been life-changing for many, so it’s no surprise that our alumni are among JAX’s most dedicated champions. We are looking forward to welcoming them back this summer as we celebrate the program’s 100th anniversary.
Grown from a modest, informal program, the SSP proudly stands today as a sought-after educational opportunity
drawing hundreds of bright applicants from all over the U.S. and beyond each year. Our students are increasingly diverse as well as talented, and they are doing work that would have been unimaginable just a few decades ago. Many will go on to careers in science and medicine, and perhaps some will even join the three SSP alumni who have gone on to become Nobel laureates. Through this change, a few things remain constant: JAX’s commitment to fostering aspiring scientists’ passion for discovery through mentored research, our transformational role in students’ lives and careers, and the importance of philanthropy in making this work possible.
The SSP centennial is not the only milestone we’re celebrating this year.
October 2024 marks the 10th anniversary of the opening of our Connecticut campus, The Jackson Laboratory for Genomic Medicine (JAX-GM). Complementing the fundamental mammalian genetics work for which our Bar Harbor campus is known,
scientists at JAX-GM are focused on human genomics. This expanded scope of our mission, in addition to fostering fruitful collaborations between our research campuses and with external partners, has created additional opportunities for SSP participants and other learners.
As meaningful as these anniversaries are, what lies ahead for JAX is even more exciting. We have ambitious plans to grow new initiatives in data science and rare disease that will amplify JAX’s impact on the scientific community and on human health. The continued support of friends and champions — like you and the alumni we are celebrating this year — are crucial to our success. As we celebrate our past and envision our future, I want to express my gratitude.

Lon Cardon, Ph.D., FMedSci President and CEO
The Jackson Laboratory


Standing the test of time
As we celebrate 100 years of JAX's Summer Student Program, we reflect on how the traditions that shape us also empower us to move forward.
by Lisa Nichols
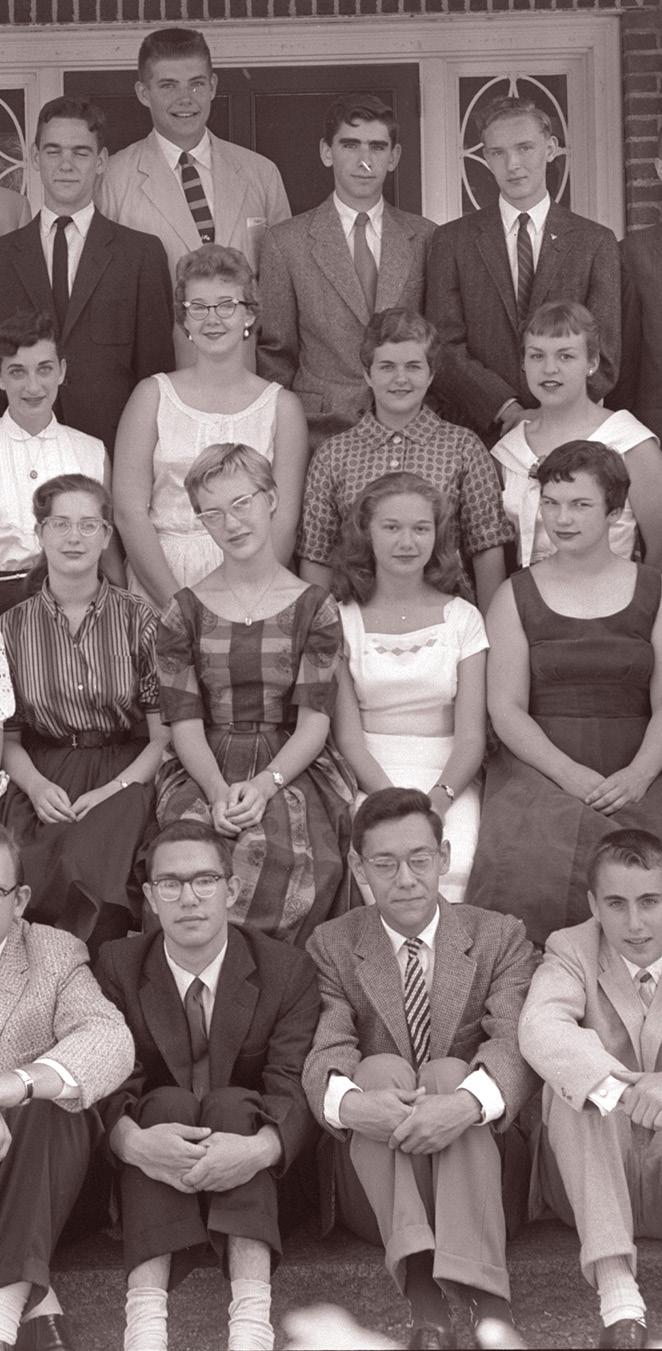
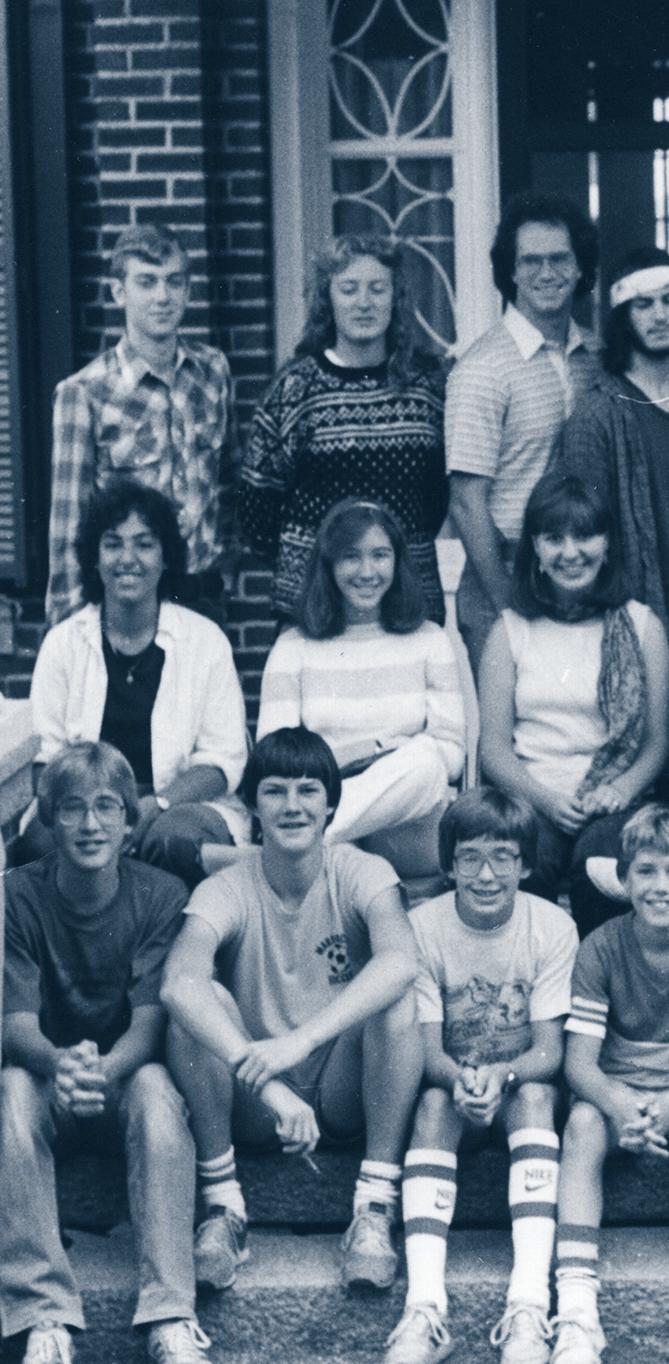


Groups of SSP participants from (l to r) 1978, 1983 and 2015
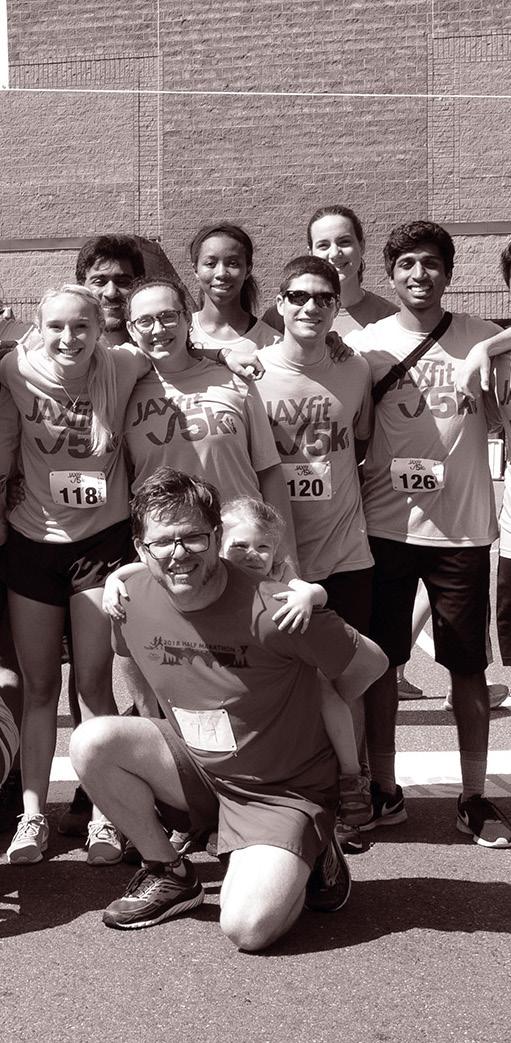

Magical. Awesome. Life-changing.
These are some of the words our alumni have used describe their experience in the Summer Student Program, which celebrates its centennial in 2024. It began with six University of Maine undergraduates who came to Maine’s Mount Desert Island for a summer of biological field studies. They slept in tents and cooked meals over open flame. Yet even in its humble early form, the program put forth a powerful hypothesis: that The Jackson Laboratory was a place where young scientists, curious in mind and adventurous in spirit, could make a difference in the world.
As all things must, the program evolved in the century since: with proper housing at the Highseas estate, for one, but also as a robust, structured 10-week program of discovery, a broader range of research options and a diverse student demographic that better reflects the world. JAX’s educational offerings have expanded to support scientists in research at every stage of their careers, including postbaccalaureate and postdoctoral talent as well as veteran experts in their field.
Yet as it always has, JAX thrums with the energy of scientists eager to make discoveries that will change our approach to the great human health challenges of our time.
This issue celebrates the qualities of JAX that endure despite change and challenge. Our tireless pursuit of groundbreaking research. Our commitment to mentorship, education and community. Our cutting-edge genomic technologies and research into complex diseases such as cancer and Alzheimer’s disease. These are the tried-and-true pillars of excellence that will propel us into the next century and beyond.
May JAX’s contributions to human health continue to be awe-inspiring, life-changing and perhaps even a bit magical, too.



Summer of discovery
by Lisa Nichols
What is the Summer Student Program?
The Jackson Laboratory’s Summer Student Program is a 10-week immersive research program for college-aged students interested in the science of genetics and genomics. Each year, approximately 40 SSP students join JAX’s campuses in Bar Harbor and Farmington for a hands-on role in our mission to make discoveries that change the future of human health. And 100 years after its founding, the program is still thriving — and still changing lives.
Humble beginnings
Since 1924, when six University of Maine students arrived in Bar Harbor for a summer of field research (the official, if informal, first cohort of SSP students), JAX’s summer student experience has been transformative for its participants. The program delivers mentorship, real-world experience and access to cutting-edge resources at a time when young minds are hungry for all three.
Based on their scientific interests, students are matched with mentors specializing in a range of research areas, including cancer, rare diseases, computational biology and neurodegenerative conditions such as Alzheimer’s disease. They work closely with a laboratory team on an independent research project and present their findings to an audience at an end-of-summer symposium.
Photo credit: Marie Chao
To date, the program has graduated more than 3,000 alumni — including three Nobel Prize winners — who have gone on to a broad range of careers in science, medicine, technology and, of course, biomedical research.
Living and learning together
Students live and work together throughout the summer as part of the SSP’s residential model.
“The fact that our students live together and learn from each other over the summer and form such close bonds makes JAX’s summer program truly unique,” says Laura Muller, director of undergraduate and STEM programs.
Students travel to and from JAX together and connect over the science, but also enjoy cooking meals together, hiking, swimming, playing games

after dinner and going to art museums. Immersed in a community that inspires them to try new experiences, SSP students find they experience countless firsttime adventures both inside and outside the lab.
Connection that endures
For many students, the SSP is just the first of many chapters in a lifelong relationship with JAX.
Some return to serve as SSP resident supervisors and staff members. Others are invited back for a consecutive summer by their mentors to continue their research together. Some alumni donate financial support to preserve the experience for future generations. And still others become a permanent part of JAX’s world-class group of scientists committed to creating a healthier future.
To learn more, visit:
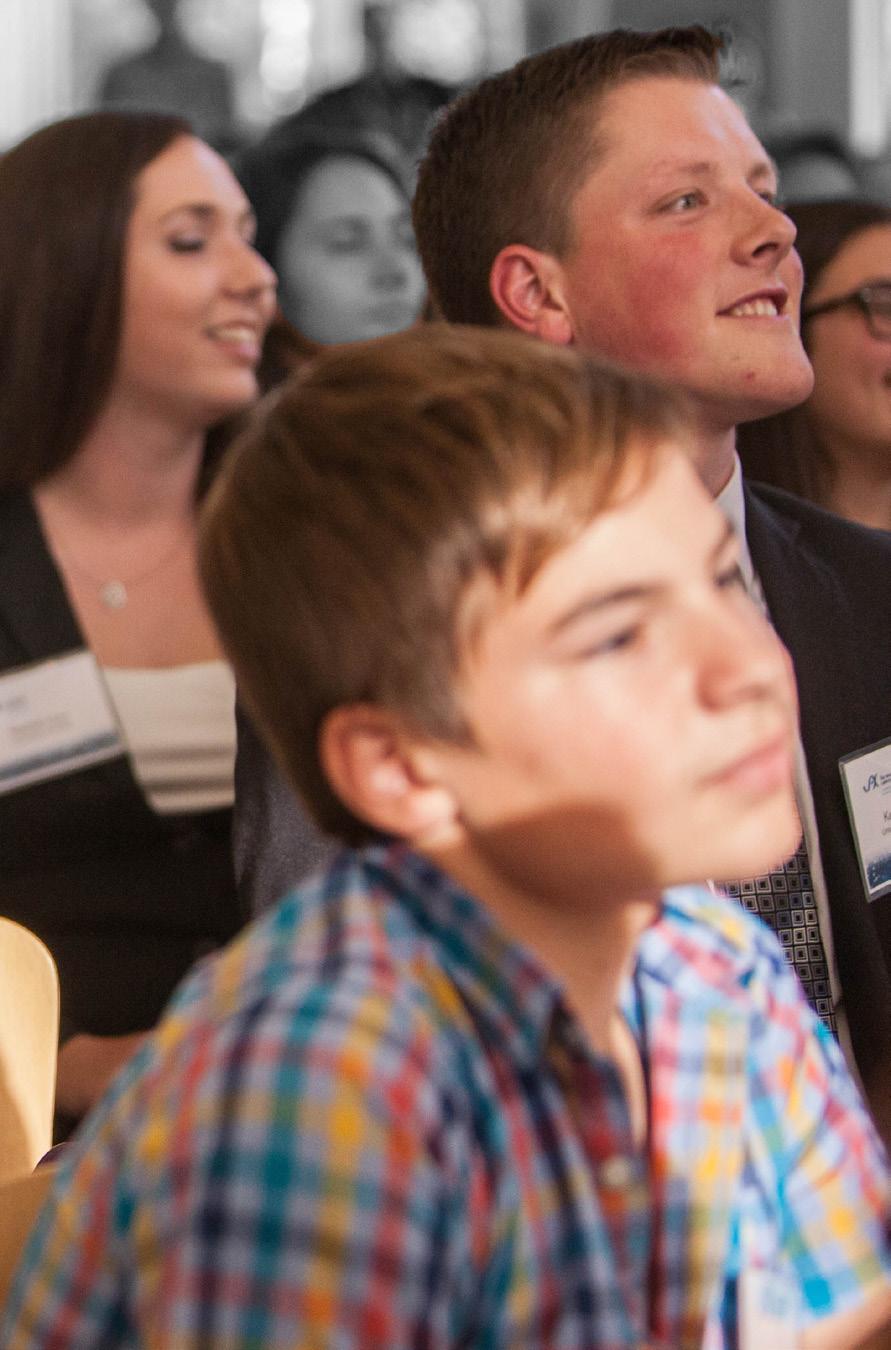


A group of SSP graduates from 2017.
Photo credit: Sarah Sullivan
The celebration of the century
By Lisa Nichols
As JAX commemorates the 100th anniversary of the Summer Student Program, we’re inviting alumni to gather in Bar Harbor, Maine to connect, celebrate and rekindle the spirit of what many still consider the best summer of their lives.









Like the simple but ambitious vision for The Jackson Laboratory’s founding — that scientists, armed with the right tools in the right environment, can make significant advances in human health — so too was the premise for the SSP’s centennial. A milestone this special calls for an unprecedented celebration.
On July 9 – 11, 2024, JAX will honor the SSP’s 100th anniversary with a special three-day program through which alumni will reconnect, experience the Laboratory in its modern-day form and immerse themselves in the splendor of JAX’s natural surroundings, as many of them did during their student days. The celebration will continue online with a series of virtual events planned throughout the summer.
“We are so excited to welcome alumni to Bar Harbor for the SSP 100th . They’ll have a chance to revisit a time that was lifechanging for so many of them and to see how far the Laboratory has come. We’re eager to share the incredibly meaningful role they have played in shaping JAX’s legacy," says AG Barry, assistant director for stewardship and alumni engagement.
Notably, Barry says, guests will also have an opportunity to spend time with current summer students at the Highseas estate, which houses students on the Bar Harbor campus.
Since 2014, the SSP has also run concurrently at the Farmington, Conn. campus where students board at the University of St. Joseph.
Summer students are a crucial part of the JAX ecosystem. They contribute to the flow of scientific ideas across the institution, amplify our reputation for groundbreaking research and affirm JAX’s enduring commitment to education.
During their 10 weeks of study, summer students benefit from attentive mentors and hands-on research that often changes the trajectory of their lives. The SSP has long been a driver of philanthropic gifts to JAX, as many alumni have been inspired to support those who follow in their footsteps.
“The shared experience of a summer at JAX is truly unique,” Barry says. “We are eager to build alumni connections across generations.”
Come join us!
The celebration coincides with the 2024 Forum for Discovery, a signature summer event that showcases faculty engaged in groundbreaking genomics research and features a special update on the Laboratory from President and CEO Lon Cardon.
Registration is now open (all guests are asked to register ahead of time). Families are welcome and encouraged to attend select events. Barry invites alumni to visit the JAX Summer Student Program Alumni Facebook page, where excitement and anticipation are already beginning to build about the prospect of a Bar Harbor summer visit. For questions and information, email AG.Barry@jax.org.
A sneak peak at the program
With alumni feedback in mind, JAX aims to evoke the most indelible student memories from inside and outside the Laboratory. Highlights include:
the “Highseas Homecoming” — an evening reception at Highseas, the early-20thcentury estate that serves as home to Bar Harbor summer students and holds generations of fond memories, where guests will enjoy light refreshments and have the chance to visit with members of the SSP class of 2024;
a volunteer service opportunity with Friends of Acadia, a leading conservation organization dedicated to the preservation and stewardship of Acadia National Park;
a special alumni on-campus networking luncheon to reunite with old friends and connect with new ones;
a walking tour of the Laboratory’s Bar Harbor campus; and
a lobster boat ride and sightseeing tour hosted on a traditional Downeast-style lobster boat.
We hope to see you there!
Five things we learned since sequencing the first human genome
by Mark Wanner


Research scientist Muneer Hasham, Ph.D.
Photo credit: Stella Fouchereaux.

The Human Genome Project wrapped up its work to sequence the first human genome in 2003 and presented it to the world in 2004. It was a true landmark achievement, but the two decades since have brought with them new insights, new understanding, new technology and new capabilities. Here are five key things scientists have learned that carry important implications for future discovery.
1. Complete doesn’t necessarily mean complete.
The original “complete” human genome sequence, as well as the vast majority of those that have followed, actually omits nearly 10% of the full sequence.
There are parts of chromosomes that are very difficult to sequence using standard methods, such as the centromeres — the middle of the “x” shape when chromosomes condense — and telomeres at the ends of each chromosome. There are also other, highly repetitive regions that are impossible to accurately align against a reference sequence, so they are omitted.
Roughly 20 years ago, when the first sequence was reported as finished, there was a lot of debate about the importance and possible function of the 98.5% of the genome that doesn’t code for proteins, so leaving certain non‑coding regions out seemed reasonable. We now know that these sequences can be quite important indeed, and recent efforts such as the T2T Consortium and the JAX led Y chromosome sequencing project have used improved long read sequencing technologies (see #4) to, at long last, fill in the gaps.
2. One genome doesn’t teach us nearly as much as we thought it would.
The first human genome sequence was generated from a sample provided by a male resident of northern European descent from the Buffalo, N.Y., region of the United States.
It was lauded as the “blueprint of life,” and it was thought at the time that it would be sufficient to provide significant insights into health and disease, even before more sequences were completed and analyzed. That has not turned out to be the case, for reasons that seem rather obvious in hindsight.
The single data set leaves out one of the two sexes entirely, of course, in addition to every other human population on earth. It’s been known for a long time that the two sexes as well as different ethnic groups carry with them different disease susceptibilities and risks, but it took a while to hammer home the message that genomic data sets need to diversify.
And while the situation has improved, most sequenced genomes are still of northern European ancestry, and much work remains to be done to make genomics more equitable.
3. Loss of function doesn’t always lead to adverse effects.
Ah, it looked so simple two decades ago. Sequence the genome, look at the coding sequences for mutations or variants associated with disease, and the roadmap to health and wellness would be clear. Well, maybe it wasn’t quite that understated, but it was close. The research since has wiped such assumptions completely away, with layers of added complexity uncovered with each new discovery.
An important finding in human datasets revealed that we all carry genetic variants that cause loss‑of‑function in many genes, with the healthy population averaging about 100 such dysfunctional genes each. Research with mice has muddied the waters further, as knocking out the same gene in multiple inbred — genetically pretty much identical — mice can result in highly variable outcomes between individuals. Sometimes it’s pretty extreme, with some mice not even surviving until birth while others live and appear pretty much normal. Therefore, interpreting genetic variation is contextual within each individual, and penetrant mutations, those that lead to consistent disease or other effects, are not the rule.



4. Most genomic variation is at the structural level.
Although human genomes are highly similar on a percentage basis, they still vary naturally at the single base level at millions of places along the ~3.2 billion base pair sequence. An adenine in my genome might be a guanine in yours, leading to the subtle changes that, in sum, make us all unique.
Little did we know 20 years ago that such differences, known to scientists as single nucleotide polymorphisms (SNPs), tell only part of the story. Other, larger variants, called structural variants (SVs) lurk, but were rarely detectable by short read sequencing methods because they don’t change a sequence that has been broken down and then reassembled. They involve sequence deletions, duplications, inversions and insertions within the genome, and they create variation for more base pairs than SNPs do.
Advanced long read sequencing methods, which can cover hundreds of thousands and even millions of bases in a single read, are now being used to detect and characterize these SVs, some of which have been associated with various diseases.

5. The road from gene to mRNA to protein is longer and twistier than thought.
While in college and graduate school too many years ago, I learned the central dogma of life. Genes, made of A, T, C and G nucleotides, are transcribed from their DNA templates to produce pre‑messenger RNAs (pre mRNAs).
The pre‑mRNAs are processed with sequences known as introns edited out. The remaining exons are stitched together into mature mRNAs. The mRNAs then travel from the nucleus to the cytoplasm, where ribosomes are located. Each three‑nucleotide sequence in the mRNA codes for a specific amino acid.
These are added one by one at the ribosome in a process called translation, until voilà, a protein is produced that will join the vast array of other proteins to carry out the functions needed by the cell. Except it’s nowhere near that simple. There is a dizzying regulatory network that determines when and how much of a gene is actually transcribed.
The processing of pre‑mRNAs is not uniform, and alternative splicing of them can create many different proteins, known as isoforms, from the same gene. Problems with any step of the process, from aberrant DNA transcription, to splicing defects, to translation misfires, can lead to dysfunction and disease.
SSP alumni share their stories


Attardi: At the time my grandmother, Tibby (Elizabeth Russell, Ph.D.), was a mouse geneticist at JAX who inspired me to apply. I majored in biology in college, then did a Ph.D. at UC Berkeley and a postdoc at MIT. I have become a mouse cancer geneticist myself, at Stanford University, where for over 20 years I have studied the p53 tumor suppressor and its roles in cancer and in developmental diseases using mouse models. JAX provided a key foundational experience for launching my career!
Van Peenan: At age 82, I still consider my weeks at JAX and Highseas to be among the most important and wonderful of my life. While all the research being conducted at the lab and its satellites is beyond important, I don’t think it is any more important than the legacy of educating and inspiring kids in the summer program.
If you had to describe your SSP experience in one word or phrase, what would it be and why?
Dubin: Mind‑blowing. I can’t think of another period since SSP in which I was exposed to so much information and so many exciting ideas.
Linda (Short) Van Peenan, SSP ’58
Kerin (Gibbons) Walsh, SSP ’83
What did you learn about yourself during your summer at JAX?
Van Peenan: That I might be a “doer” rather than a “thinker.” This became more and more apparent as I progressed through school and became a surgeon rather than a research scientist.
Isola: I learned that I was normal — or at least that there were other teens like me — so I felt like I found a place that I belonged. That gave me more confidence that I could just be myself.
Walsh: Since this was the summer I graduated from college, I developed a greater confidence in myself and a belief that I could achieve my career goals.
How
did the Summer Student Program shape your career or your post-JAX plans?
Van Peenan: This program was the first “credential” I earned. It looked wonderful on an application for a Fleming Scholarship at Oklahoma Medical Research Foundation the following summer. These two credentials mushroomed and eventually led to my acceptance to medical school and a vascular surgery fellowship. Bottom line: My summer at JAX started it all.
Mary E. Delany, SSP 78': I decided to pursue a career in research and in genetics and specializing in cytogenetics. With (JAX Professor Emeritus) Muriel Davisson’s recommendation, I landed a research technician position with her former Ph.D. mentor at Penn State. From there I went to Cornell University for M.S. and Ph.D. degrees and ultimately became a professor at UC Davis.



Vicki (Salonen) Isola, SSP ’77
Steve Dubin, SSP ’77
Laura Attardi, SSP ’86
100 years on
Preserving the Summer Student Program’s essence of professional and scientific exploration with Program
Director Laura Muller, Ph.D.
by Sophia Anderson
In a century marked by rapid scientific and technological advancement, the Summer Student Program has remained steadfast in its mission to create a focused, exploratory experience for young minds interested in research.
Originating in the forests of Acadia, the SSP has since helped to forge leaders and innovators in both scientific and non-scientific careers, and with the celebration of the program’s 100th year, it is only fitting to showcase its essential tenets. Explore the program’s legacy and goals alongside Muller as she details the core values of the program and why they have contributed to its success.
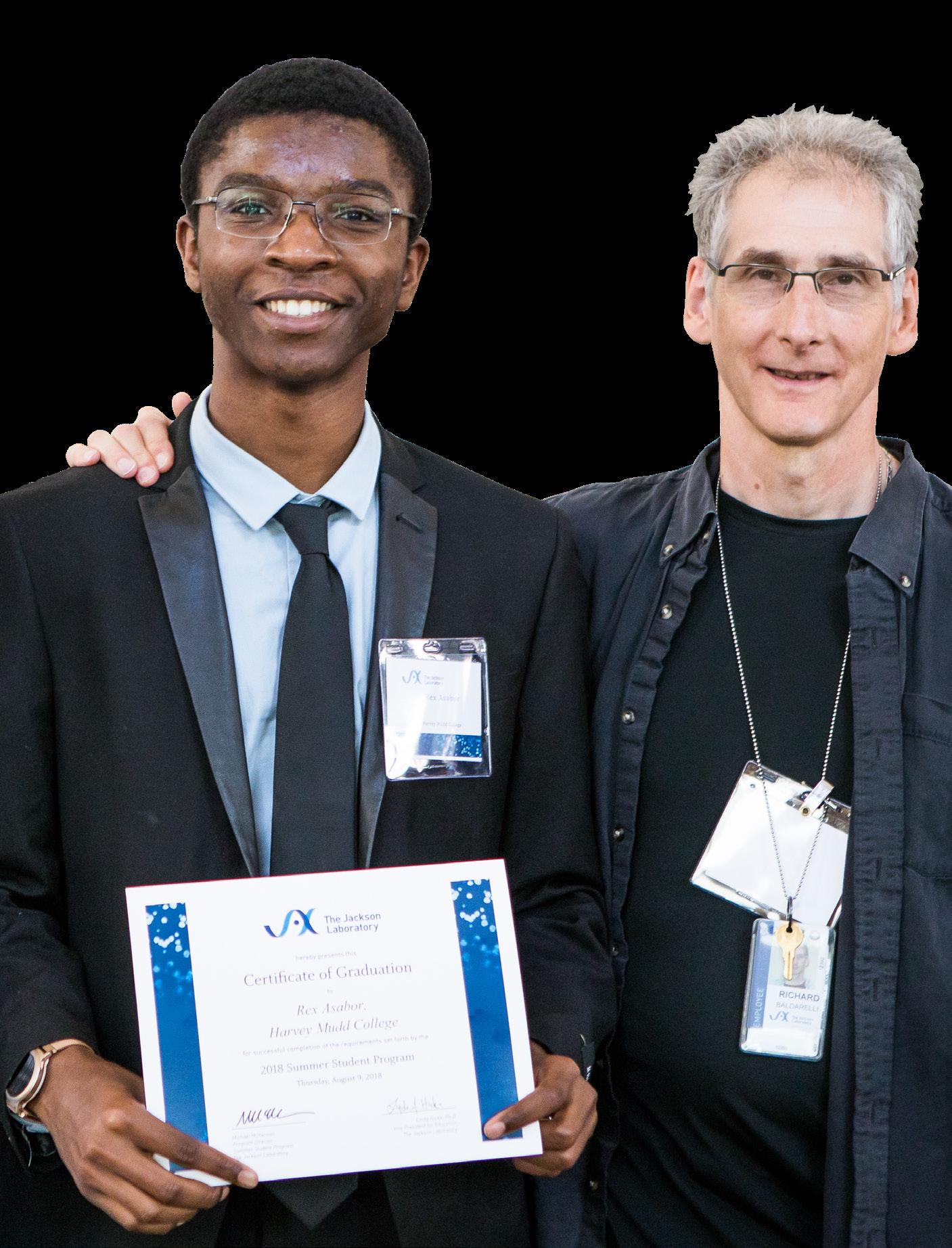





(l to r) Senior research scientist Joel Richardson, Ph.D.; Summer Student Rex Asabor; and Bioinformatics
Analyst Richard Baldarelli, Ph.D., pose during the 2018 SSP symposium. Photo credit: Tiffany Laufer.
Immersive
One of the reasons why the SSP has been successful is its commitment to curating a welcoming environment for the next generation of brilliant minds. Community and belonging are major cornerstones of the SSP experience and are instilled into the structure of the program from day one.
Throughout their time in either Bar Harbor, Maine or Farmington, Conn., students are completely immersed in a research environment alongside their peers. Surrounded by people who have similar interests and are around their own age, individuals develop friendships within a culture of camaraderie despite being far from home.
“The SSP provides the answer to the question, 'What does it mean to be a professional scientist?'
By being away from their typical environment, students are able to experience how it feels to be in a community of like-minded people and can begin to picture where they could contribute," says Muller.
Each SSP student delves headfirst into their research area of interest, exploring their new ecosystem and engaging in collaborative projects. The importance of connection in the scientific field is reinforced over the course of the summer, as they engage with lab members, faculty and staff at The Jackson Laboratory.
By instilling the importance of connection into the program, JAX encourages students to be engaged participants in whatever they do during and well after they finish their education and training.
Inclusive
From sea to shining sea, the SSP has reached students across the United States. Over the last 100 years, representatives from all 50 states have attended the program, including those from underrepresented or disadvantaged backgrounds. Based on criteria from the National Institutes of Health, 59% of students enrolled in the SSP were from underrepresented groups over the past five years.
“It is an important fact about who we are,” says Muller. “We are able to bring a significant number of students who, based on their background, would not have been able to do this kind of research.”
Showcasing the importance of diversity in shaping the scientific landscape, the SSP is not solely about academic pursuits. Part of what makes the program unique is the way it provides for its accepted students. In addition to flights, food and housing being provided, each student receives a stipend, so they end the summer with not only a one-of-a-kind experience under their belts but also money in their pockets.
“We really try to pick students with promise. Students can be limited by their zip code or by what their high school has to offer for science courses. They may still be first year students in college who have not had the opportunity to get in the lab yet. We accept them into our program, they do research for the first time and they catch the bug,” says Muller.
Innovative
Thinking outside the box often leads to the greatest breakthroughs, especially in science. The SSP encourages individuals to express their inventiveness by incorporating a hands-on, experiential learning approach. Many accepted students have had little to no opportunity to perform in-depth benchwork prior to their enrollment. The program provides novice yet promising students with the chance to develop laboratory skills in state-of-the-art facilities, supporting their budding critical thinking capabilities.
“We select students who show a real interest in science and understand what research is about. Maybe they have not done any formal research projects, but they can recognize the times in their lives when they have performed their own type of research,” says Muller.
Whether it’s testing a new recipe in the kitchen or attempting a technique during ceramics class, the way students perform research in their daily lives takes many forms. The SSP reinforces the observant and resourceful nature of these students by giving them an opportunity to learn both hard and soft skills that they will carry into whatever career they choose.

Impactful
All scientists understand that their work stands on the shoulders of those who came before. The same can be said for the SSP. At the end of each summer, each cohort joins a network of alumni who will forge ahead with research — or other fields — as they make their way into the real world.
The impact of the program has ripple effects throughout numerous fields, and alumni have gone on to start research labs, become medical practitioners, advocate for research funding on Capitol Hill, educate other students of promise and more.
Many remain connected and continue to engage with the program long after their summer of research has ended, which speaks volumes about how much they value this experience.
The spirit of exploration, innovation and inclusivity that defines the SSP is encapsulated here as a testament to its accomplishments over the past 100 years. These principles will be a reminder to future students and educators, illustrating how far the SSP has come and its commitment to educating forward thinkers in the ever-evolving landscape of scientific discovery.

The evolution of a career in research
by Mark Wanner
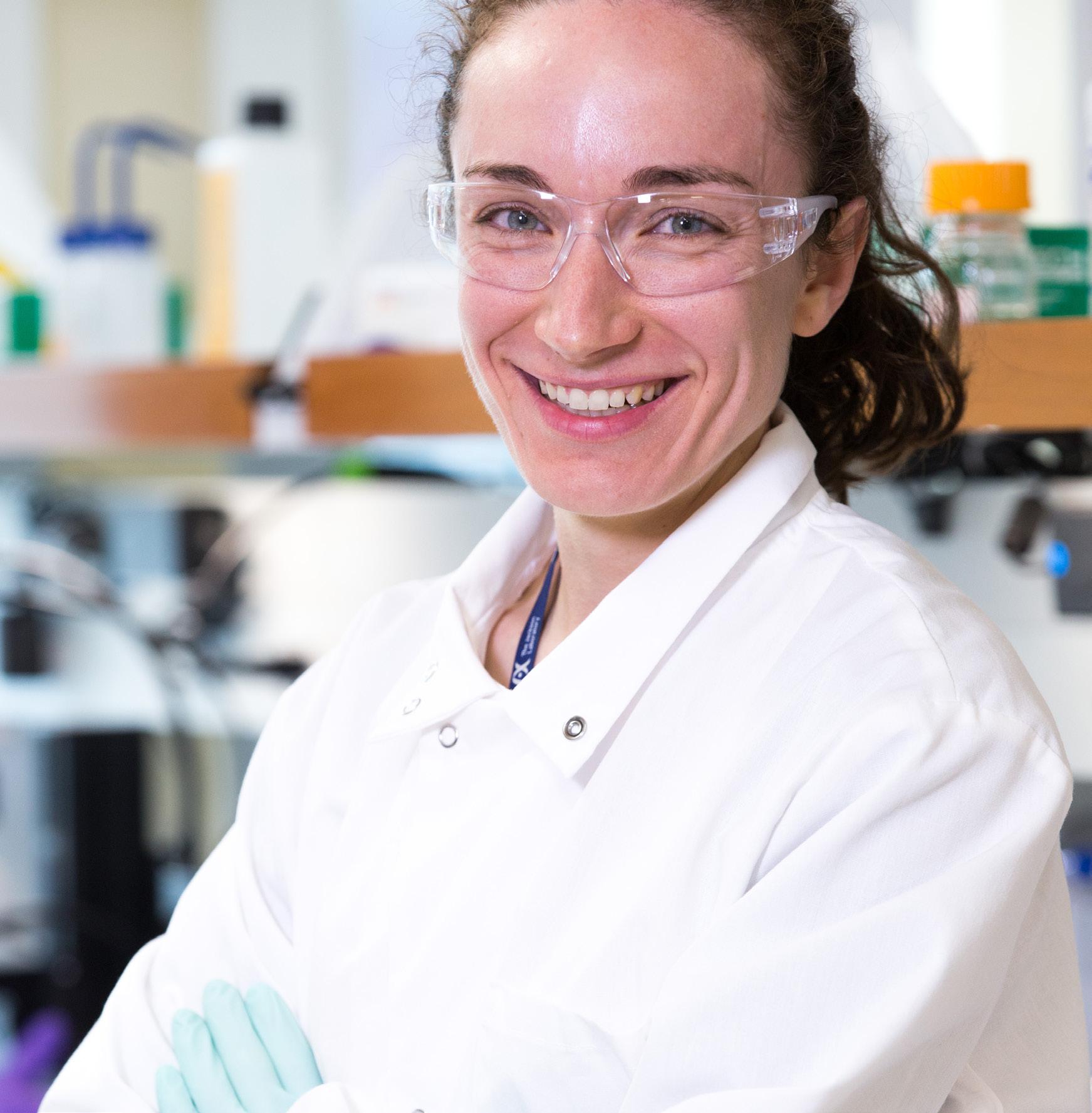




It was the summer of 2002. “Hot in Herre” by Nelly topped the music charts. The Boston Red Sox were toiling through year 84 of the Curse of the Bambino. And, for the 78th year, students gathered in Bar Harbor, Maine, for an immersive research experience. It was an intense experience for all, but, for some, it would be life-changing.
Among those who traveled to The Jackson Laboratory campus in Bar Harbor for the renowned Summer Student Program that year was Beth Dumont, a rising sophomore at Cornell University. A native of Augusta, Maine, she had wanted to spend the summer closer to her family. She came across the SSP somewhat by accident, but, as a biology major, it seemed like an interesting way to return to her home state for the summer.
“I’d heard of JAX but only as a mouse supplier, so I had no idea what I was getting myself into,” says Dumont.
“I didn’t know about research outside of basic freshman biology and chemistry labs. But it was awesome.”
Fast forward more than two decades, and Dumont is an associate professor on the JAX faculty, a leader in the field of population genetics and a mentor for students each summer. Her experience bridges JAX’s past and present and highlights what makes it so unique and so important in the research world.
Associate Professor Beth Dumont, Ph.D.
Photo credit: Tiffany Laufer.
The wonders of variation
Spending her summer in the lab of Beverly Paigen, Ph.D., Dumont was exposed to a research program covering wideranging interests. When she first arrived, she had no experience with even the most basic lab techniques, including how to pipette. Paigen assigned Dumont a postdoctoral associate to supervise her directly, but he was at a conference when she first stepped into the lab. In stepped another postdoc, Ron Korstanje, Ph.D., to get her off to a good start. As fate would have it, Korstanje would also subsequently join the JAX faculty and work with Dumont as a colleague. And to truly bring things full circle, Maike Korstanje, Ron’s daughter, recently completed an academic year internship in the Dumont lab.
Although she didn’t know it at the time, Korstanje’s guidance helped launch Dumont on a project that would shape her future training and career. Her work that summer contributed to a project that aimed to identify genomic regions contributing to differences in susceptibility to cholesterol gallstone disease between mouse strains. Mapping studies in the early 2000s rarely succeeded in identifying single genes contributing to disease, but rather localized genetic effects on a trait of interest to broad regions of the genome — termed loci — containing multiple genes. Dumont’s effort focused on identifying compelling candidate genes within these mapped loci using the nascent mouse genome sequence and testing candidate genes for sequence and expression differences between strains.
“With the project, I grew to appreciate the ways we can leverage the differences between mouse strains to pinpoint the genetics that underlie disease susceptibilities,” says Dumont. “But this led to other questions: How and why do the differences arise in the first place? And what roles do they play in function and biology?”
The fundamentals
The answers to those questions lie in the field of population genetics, which Dumont characterizes as the study of where genetic variation comes from and why it exists in the patterns that it does through time and space. In humans, it’s vital for studying how population differences contribute to variable disease susceptibility and risk. Dumont returned to Cornell enthusiastic and motivated to pursue this new-to-her research area, and she reached out to faculty working in the field. One needed help, and he subsequently hired her to work in his lab as an undergraduate. Fortuitously, Dumont had connected with Andrew Clark, Ph.D., someone who was not exactly a typical professor and also had ties with JAX.
“Andy had been involved with the Human Genome Project and literally wrote the book on population genetics, and I’m glad I didn’t know beforehand that he was such a bigshot in the field!” says Dumont. “He was also a regular instructor at the McKusick Short Course, so he knew JAX as well. I worked with him throughout my remaining time at Cornell.”

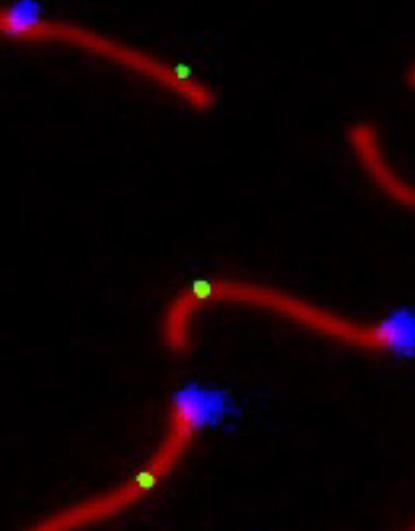
Working with Clark provided a further boost to Dumont’s studies when she went to graduate school and did her thesis work with a former postdoc of Clark’s at the University of Wisconsin-Madison. There she focused on the genetic control of recombination rates. Recombination is a fundamental but somewhat counterintuitive biological mechanism, in that it involves breaking the genome in many places and then repairing the breaks, resulting in a scrambling of genetic traits for the next generation. It is a necessary step at the onset of meiosis, in which mature gametes (eggs or sperm) are produced, each ultimately containing only one set of chromosomes rather than the two present in other cells.
Evolution in the lab
Ph.D. in hand, Dumont went on to postdoctoral appointments at the University of Washington and then North Carolina State University.
At NCSU she was awarded a federal grant, called a K99, to enhance her independent research. Her work expanded her exploration of recombination biology,
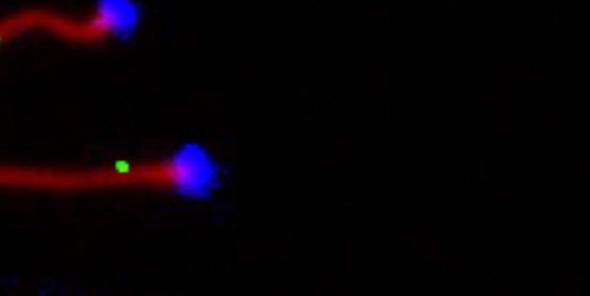
honing in on sex chromosomes and the regions on them essential for recombination and proper segregation during meiosis. It also provided her with the experience she needed for her next step forward: starting her own independent laboratory. And in 2016, she returned to JAX to do just that.
Now Dumont works with a unique resource at JAX: diverse mouse populations. Her work still investigates recombination and other biological mechanisms, but she is now seeking to answer even larger questions: How do species originate and evolve?
The genetically diverse mice provide her with essential insights.
“I’m
thrilled to be at JAX, because many of the projects using genetically diverse mice are basically beautifully curated evolutionary experiments in action,” says Dumont. “I
can track the
process
of
DNA transmission
from one
generation
to the next because we know how the DNA segregates between generations. We can learn how many mutations accumulate, how and where they accumulate … it’s evolution in a self-contained environment.”
She also gives back, seeking to work with students each summer who, much like herself when she first arrived at JAX, may not have previously thought about research as a career.
“It’s wonderful to be on the other side and to be able to nurture the potential in students over the course of the 10-week program,” she says. “I also keep in touch with (now) colleagues who went through the program and follow their research trajectories. For many, like me, it was a transformative experience and provided opportunities that they otherwise wouldn’t have had.”
Chromosomes during meiosis, with the sites of crossover repair indicated by green dots. Photo credit: Beth Dumont.
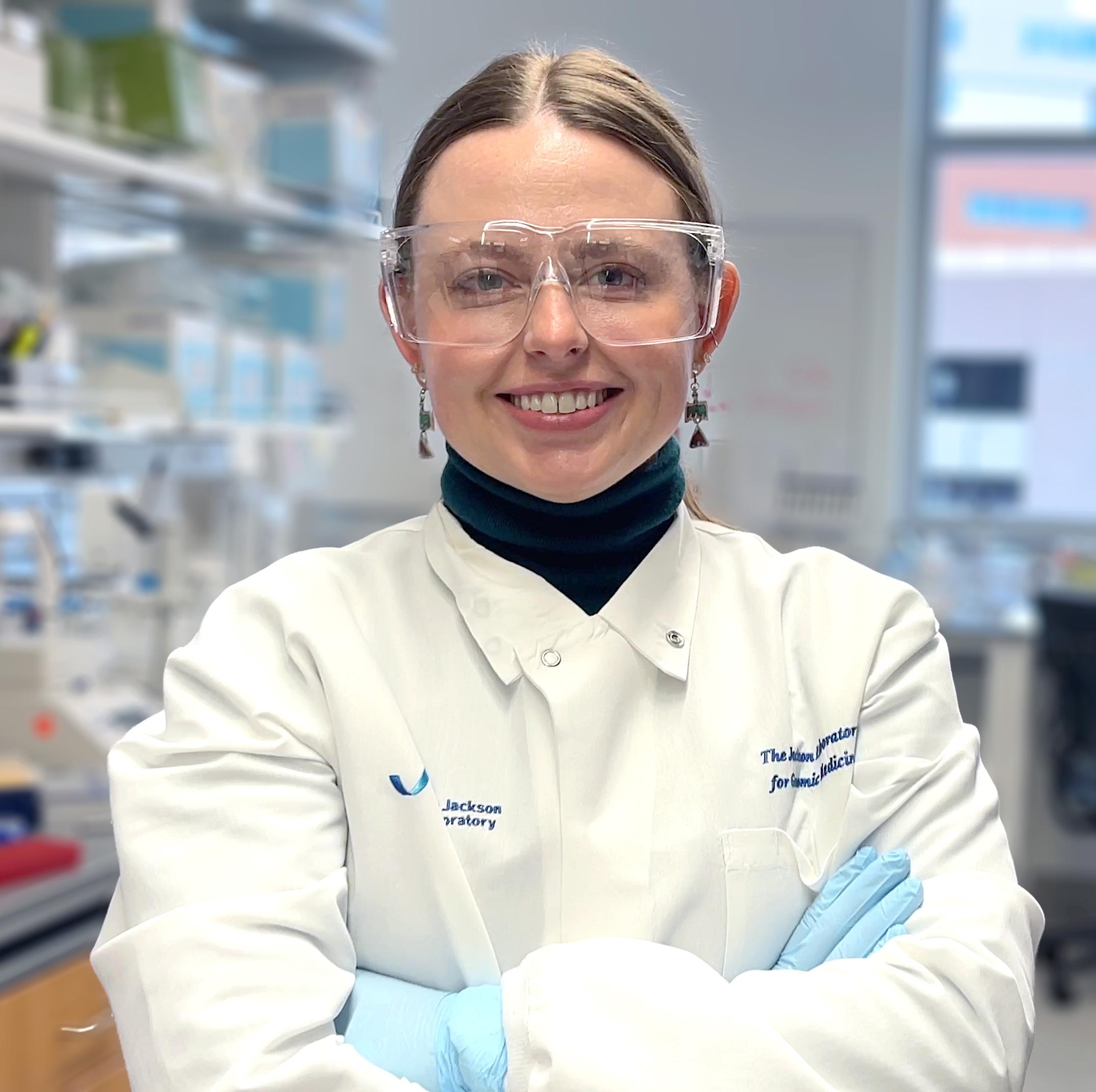




Morgan Severn
Photo credit: Sophia Anderson.
Takeshi Iwasaki
Photo credit: Takeshi Iwasaki
The JAX Scholars Program supports exceptional postdoctoral associates as they pursue better understanding of mammalian biology and human disease. The Jackson Laboratory is proud to announce Morgan Severn, Ph.D., and Takeshi Iwasaki, M.D., Ph.D., as the 2024 JAX Scholars.
Budding experts in the fields of immunology and microbiome named as 2024 JAX Scholars
by Sophia Anderson
“We are very proud of our JAX Scholars. They are wonderful examples of our diverse community of postdocs who creatively apply their expertise to uncover the genetic underpinnings of human health. Iwasaki and Severn exemplify the value in assembling an interdisciplinary group of scientists with different academic backgrounds who can leverage their collective knowledge toward successful research outcomes,” says Meredith Theeman, Ph.D., director of predoctoral and postdoctoral education.
The variant investigator
With a wealth of experience in the clinical field, Takeshi Iwasaki is determined to have a positive impact on patient outcomes. During his residency, Iwasaki managed multiple cases where patients’ diagnoses led to a dead end, with too little known about their specific condition to provide them with targeted treatments. Despite the tremendous amount of progress over the past decade to identify genes and regulatory elements in the human genome, cases like those Iwasaki encountered are still far too common, and there remains a pressing need to link genetic variants to disease presentation or risk. Identifying the actual genetic culprits amid the vast number of potential disease-contributing factors is a difficult task, but Iwasaki is up for the challenge as he pursues his career as a physician-scientist.

Iwasaki’s graduate research focused on complex autoimmune diseases, including the study of rheumatoid arthritis patients’ responses to tumor necrosis factor inhibitor (TNFi), one of the key drugs in treating the disease.
His findings uncovered a systemic pattern of events occurring in TNFi non-responders. Using this information, he was able to create a tool to predict when the treatment would be ineffective for these patients. Iwasaki also profiled 581 clinical presentations for systemic lupus erythematosus, better known as lupus. He uncovered 26 correlations among clinical sub-phenotypes of the disease and created an accurate statistical model to calculate the probability of exhibiting 10 out of 20 phenotypes. Much of this work hinged on identifying what genomic variants were associated with these autoimmune diseases, but findings in the transcriptome, the collection of all RNA or gene readouts, left Iwasaki with the desire to know more.
“When
I observed how the transcriptome is elevated with specific diseases, I realized I could not prove the causality of this phenomenon,” says Iwasaki. “This drove me to pursue a postdoctoral position to acquire specific experimental techniques to prove what was causing these changes.”

Photo credit: Cloe Poisson
Iwasaki will be joining JAX Associate Professor Ryan Tewhey, Ph.D., as a postdoctoral associate this spring. He will be continuing his work identifying variants responsible for autoimmune disease onset by establishing a platform that uses cutting-edge assays to understand how variants in the genome impact disease risk, including high-throughput CRISPR screens. Unraveling the effects of genomic variants will help Iwasaki and colleagues in the Tewhey lab determine the precise genetic mechanisms influencing autoimmune disease, potentially paving the way for novel therapeutic approaches.

“We are delighted to have Iwasaki join our group. His clinical experience and expertise in human genetics will be a perfect complement to the diverse group of engineers, biologists, geneticists and computational scientists in the Tewhey Lab. I am excited to work with Iwasaki to apply the tools we develop to investigate the genetic causes for autoimmune disorders,” says Tewhey.



“I am honored to receive the JAX Scholar award. This is the first time I have received this kind of support,” says Iwasaki. “I am excited to work with Tewhey as our interests and current research align. With my background as an M.D. and computational skills, I believe I will be able to meaningfully contribute to the scientific world.”
Photo credit:
Takeshi
Iwasaki
The microbe whisperer
Driven by her curiosity, Morgan Severn is committed to making an impact on human health. Although initially unsure of her career path, Severn identified her passion for research after an undergraduate internship in a microbiology laboratory, where she realized there was still much to uncover about how we interact with the billions of microbes living in and on us.
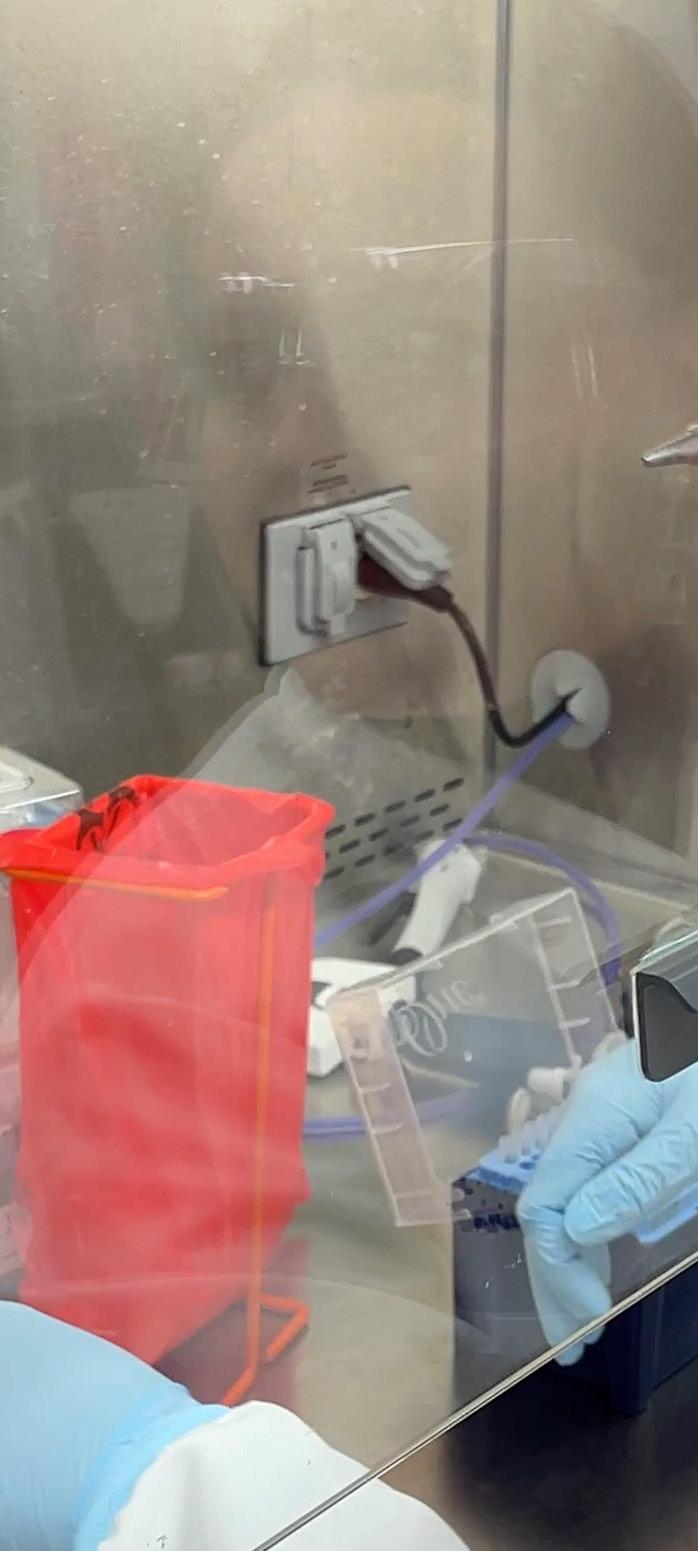


During her graduate studies, Severn explored microbial quorum sensing, also known as microbial communication, with a focus on how common members of the skin microbiota interact. She was most interested in uncovering the communication strategies of the understudied types of staphylococci naturally residing on the skin. One of Severn’s key findings highlighted that S. aureus, a common and widespread bacterial pathogen known to cause severe skin and soft tissue infections, quorum sensing could be blocked
“When
I found microbiology, it felt like a natural fit for me,” says Severn. “I love being around people who are inquisitive and excited about asking interesting questions related to microbial physiology and the microbiome.”
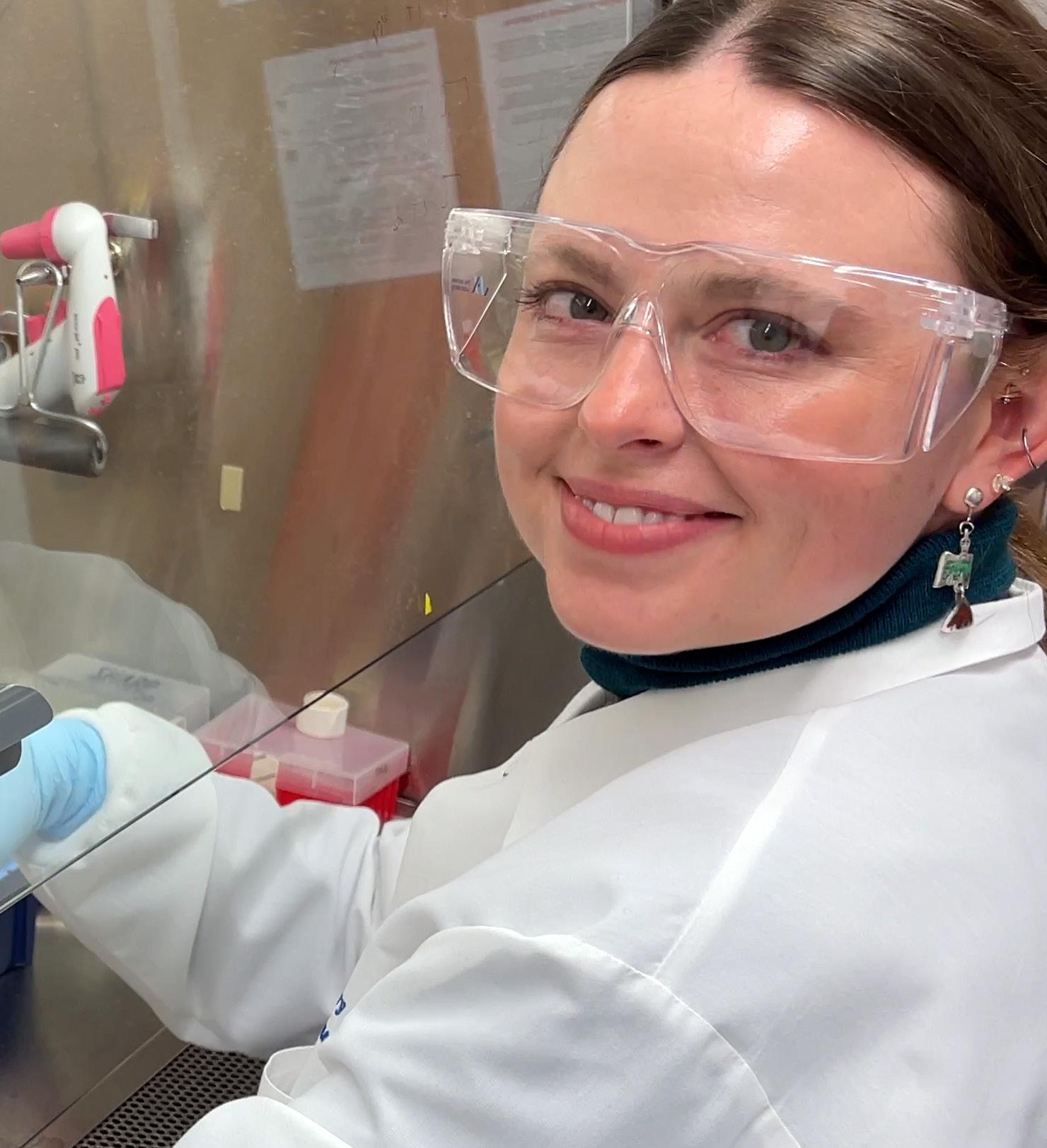

by another ubiquitous colonizer of the skin, S. hominis, which prevented S. aureus from causing infections. Now as part of the lab of JAX Associate Professor Julia Oh, Ph.D., Severn studies staphylococci as well as another common member of the skin microbiota known as corynebacteria. Despite corynebacteria being some of the most abundant bacteria on the skin, little is known about how they interact with their human hosts. Severn hypothesizes that they likely play an important role in maintaining balance on and within the skin.
Photo credit: Stella Fouchereaux
“Severn is an outstanding expert in staphylococcal biology. It will be so exciting to see how she will continue to integrate her classical microbiology training with cuttingedge ‛omics approaches to blow open the field’s understanding of host–microbiome interactions in the skin,” says Oh.



“We are currently growing corynebacteria on human skin organoids to see how these tissues respond. We have observed a distinct category of bacteria that do not stimulate an immune response, while another subset of the bacteria is much more pro-inflammatory. Now we are trying to understand and dissect what makes these two groups unique from each other and between strains,” says Severn.
With further study, Severn believes there is potential to use skin microbes like corynes or staphy as biotherapeutics to treat common skin diseases such as eczema or acne. By taking members of an individual’s own microbial community, genetically repurposing them and then reapplying them to the skin, it may one day be possible to treat these common dermatologic problems. Severn is grateful for the unwavering support and mentorship provided by Oh and is confident that working at JAX will prepare her for running her own lab one day.
“JAX is a supportive and inclusive place to do great science,” says Severn. “Oh has been a fantastic mentor and always wants to help us utilize the services and resources available. The support for trainees here allows us more opportunities to ask and answer important biological questions.”
Photo credit: Stella Fouchereaux


