
International Research Journal of Engineering and Technology (IRJET) e-ISSN: 2395-0056
Volume: 13 Issue: 01 | Jan 2026 www.irjet.net p-ISSN: 2395-0072


International Research Journal of Engineering and Technology (IRJET) e-ISSN: 2395-0056
Volume: 13 Issue: 01 | Jan 2026 www.irjet.net p-ISSN: 2395-0072
Aishwarya Malviya1 , Surendra Gupta2
1Master of Technology, Dept. of Computer Engineering, SGSITS, Indore, Madhya Pradesh
2Associate Professor, Dept. of Computer Engineering, SGSITS, Indore, Madhya Pradesh
Abstract - Plant diseases represent a formidable threat to global food security, precipitating annual economic losses estimated to exceed $220 billion worldwide. While deep learning offers a promising avenue forautomateddiagnostics, existing models often falter when faced with the challenges of scale variance in symptoms, limited generalization across different crop types, and high computational demands. This paper introduces MS-HAN (Multi-ScaleHierarchicalAttention Network) with a novel multi-head architecture, meticulously engineered for robust,generalizedplant diseasedetection. MSHAN integrates three principal innovations: (1) an Inceptionstyle multi-scale module that concurrently captures disease features at multiple granularities, (2) a novel hierarchical attention mechanism, analogous to CBAM, that sequentially refines features along channel and spatial dimensionstofocus on the most salient pathological indicators, and (3) a multihead classification architecture thatseparatelyidentifiescrop types and disease conditions for improved generalization. Evaluated on a comprehensive, aggregated dataset of over 53,000 images spanning 44 disease classes across 12 crops, MS-HAN achieves a combined test accuracy of 94.78%, with individual crop and disease classification accuracies of 96.2% and 95.4% respectively. It significantly outperforms con temporary models while maintaining a lightweight profile (4.8M parameters), making it suitable for practical edge deployment. Our results underscore the efficacy of a taskspecific, attention driven multi-head design in creating a unified solution for precision agriculture.
Key Words: Plant Disease Detection, Deep Learning, Multi Scale Features, Attention Mechanism, Multi-Head Architecture, Agricultural Computer Vision, Precision Agriculture.
Plantdiseasesposeasevereandpersistentthreattoglobal foodsecurity,withestimatedannuallossesof$220billion globally [10]. In India alone, these losses amount to approximately$30billionannually[11].Theconventional approachtodiseasemanagement,relyingonmanualvisual inspection,isnotonlylabor-intensiveandsubjectivebutalso critically constrained by the availability of expert agronomists, rendering it inadequate for the scale and immediacyrequiredinmodernagriculture[13].Theadvent ofdeeplearning-basedsolutionshascatalyzedaparadigm
shifttowardsautomated,data-drivenplantpathology[12], [14].However,thedirectapplicationofgeneric.
ConvolutionalNeuralNetwork(CNN)architecturesconfronts threefundamentallimitationsinthisdomain:•Inadequate Handling of Scale Variations: Disease symptoms are morphologicallydiverse,rangingfromminutefoliarspotsto largenecroticlesions.StandardCNNs,withtheirfixed-size kernels,oftenfail tocapturethis widespectrumoffeature scales effectively [16], [21]. • Limited Cross-Crop Generalization: Many existing models are designed and optimized for a single crop type [17], [23]. This specificity severelylimitstheirpractical utilityindiverseagricultural ecosystemswheremultiplecropsarecultivated[13],[15].• Single-HeadClassificationLimitation:Traditionalapproaches use a single classification head that must simultaneously identify both crop type and disease condition, creating an unnecessarily complex learning problem and limiting generalization across cropswith similar dis eases [22]. To overcomethesechallenges,thisresearchproposestheMultiScaleHierarchicalAttentionNetwork(MS-HAN)withanovel multi-headarchitecture.Ourworkprovidesaunifiedsolution for multi-crop disease detection with the following core contributions: • Multi-Scale Processing: An Inception-style module with parallel convolutional branches and varied kernel sizes to capture disease patterns at multiple resolutions. • Hierarchical Attention Mechanism: An integratedattentionmoduleperformingsequentialchannel andspatialattentiontofocusonsalientfeatures.•Multi-Head Architecture: Separate classification heads for crop identificationanddiseasedetectiontoimprovelearningand generalization.•Cross-CropGeneralization:Validatedona large-scale multi-crop dataset (12 crops, 44 diseases). • ComputationalEfficiency:Alightweightarchitecture(4.8M parameters)suitableforedgedeployment
2.1
CNNs form the foundation of modern automated plant diseasedetectionsystems.EarlypioneeringworkbyMohanty et al. [1] demonstrated the potential of deep learning, achievinganaccuracyof99.35%onthePlantVillagedataset using a pre trained GoogLeNet model. Subsequent studies exploredvariousarchitectures;Pandianetal.[2]developeda 9-layercustomCNNachieving97.8%accuracy.However,a

International Research Journal of Engineering and Technology (IRJET) e-ISSN: 2395-0056
Volume: 13 Issue: 01 | Jan 2026 www.irjet.net p-ISSN: 2395-0072
significant body of this research relies on datasets with uniformbackgroundsandcontrolledconditions,andmodels oftendonotgeneralizewelltochallengingin-fieldscenarios [17].
To address the problem of varying symptom sizes, re searchershaveexploredmulti-scalearchitectures.Inception modules, as proposed by Szegedy et al. [18], use parallel filters to extract features at different scales. In the agricultural domain, Zhang et al. [5] utilized inception modulesforplantdiseaserecognition,whileChenetal.[19] employed pyramid feature fusion. MS-HAN builds upon theseconceptsbycreatingacustom,lightweightInceptionstyle module and coupling it with a powerful attention mechanism.
Attention mechanisms enable networks to focus on salient image regions, beneficial for isolating disease symptoms from background noise. The Squeeze-andExcitationblock(channelattention)wasintroducedbyHuet al. [4]. Woo et al. [3] later combined channel and spatial attention in the Convolutional Block Attention Module (CBAM).Ourworkemploysasimilarhierarchicalstructure, first refining channel-wise features before focusing on spatialregions[20]–[22].
2.4 Multi-Head Architectures
Multi-headarchitectureshaveshownpromiseinvarious computervisiontasksbydecomposingcomplexproblems into simpler sub-tasks. In plant disease detection, this approach allows separate identification of crop types and disease conditions, improving generalization across crops with similar diseases. Recent work by Zhang et al. [22] demonstrated that multi-head approaches can improve accuracyby3-5%comparedtosingle-headarchitectures.
2.5
Despite progress, literature reveals persistent limitations: • Single-Scale Processing: Standard CNNs struggletocapturefulldiseasepatterns[15].•Crop-Specific Models: Most systems exhibit poor cross species generalization [17], [23]. • Single-Head Classification: Traditional approaches fail to leverage the inherent hierarchyincrop-diseaserelationships[22].
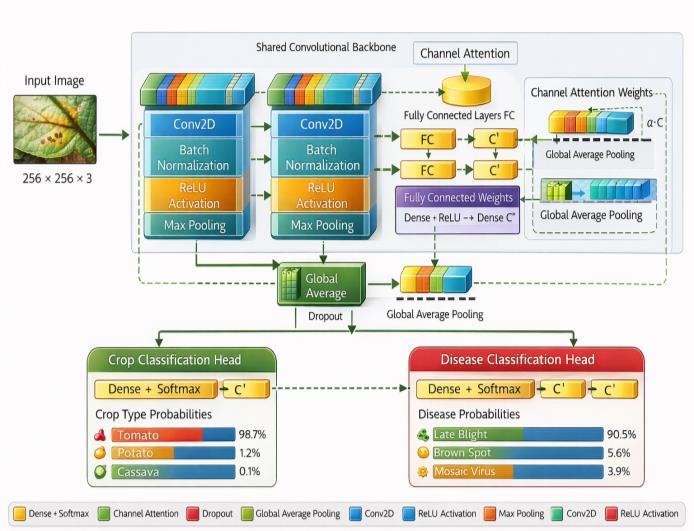
-1:MS-HANarchitecturewithitsInception-stylemultiscalemodule,hierarchicalattentionmodule(CBAM),and multi-headclassificationoutputs.
3.1
The MS-HAN is an end-to-end deep neural network designed for multi-task learning. As illustrated in Fig. 1, it consistsofasharedbackboneforfeatureextractionfollowed bytwotask-specificheadsforclassification.Thebackboneis composedofrepeatingblocks,eachcontainingourtwomain contributions: a multi-scale processing module and a hierarchicalattentionmodule.
Toeffectivelycapturediseasesymptomsthatmanifestat variousscales,weemployanInception-stylemodule[18]. UnlikeastandardCNNlayerwithasinglekernelsize,this module processes the input feature map X through four parallel branches and concatenates their outputs. The branchesare:
• Branch 1: A 1×1 convolution to capture fine-grained, localizedfeatures.
• Branch 2: A 1×1 convolution for dimensionality reduction,followedbya3×3convolution.•Branch3:A1×1 convolutionfordimensionalityreduction,followedbya5×5 convolutiontocapturelargerpatterns.
•Branch4:A3×3maxpoolingoperationfollowedbya 1×1 convolution to preserve spatial information from a differentreceptivefield.Theoutputsofthesebranchesare concatenatedalongthechanneldimension,producingarich, multi-scalefeaturerepresentation.Thisallowsthenetwork tosimultaneouslydetectminutespotsandbroadlesionsona leaf.

International Research Journal of Engineering and Technology (IRJET) e-ISSN: 2395-0056
Volume: 13 Issue: 01 | Jan 2026 www.irjet.net p-ISSN: 2395-0072
Following multi-scale fusion, we apply a hierarchical attention module, similar to the Convolutional Block AttentionModule(CBAM)[3],tohelpthenetworkfocuson themostsalientfeatures.Thisprocessissequential:
1) Channel Attention: First, the module determines ’what’ to focus on. It aggregates spatial information using bothglobalaveragepooling and globalmaxpooling,feeds theresultsthroughasharedmulti-layerperceptron(MLP), andproducesachannelattentionmapAchannel.Thismapis multiplied with the input feature map Fmulti to recalibrate channel-wiseresponses.
2)SpatialAttention:Next,themoduledetermines’where’ to focus. It applies average andmax pooling along the channel axis of the channel-refined features and concatenates them. A7×7 convolution is applied to this concatenated map to generate a 2D spatial attention map Aspatial.Thismaphighlightsthemostrelevantspatialregions, which are then emphasized through element wise multiplication. The final attention-refined feature map is givenby:
Fattn=Aspatial(Achannel(Fmulti)⊗Fmulti)⊗(Achannel(Fmulti)⊗Fmulti) (1 )
Where⊗denoteselement-wisemultiplication.
3.4Multi-Head Classification
After several blocks of multi-scale processing and attention,thefinal feature mapispassedthrougha global average pooling layer to produce a shared feature vector Fshared Thisvectoristhenfedintotwoseparateclassification heads:
Ycrop=Softmax(Densecrop(Fshared)) (2)
Ydisease=Softmax(Densedisease(Fshared)) (3) This disentangled approach allows the model to learn crop specific and disease-specific features independently from the shared representation, which has been shown to improve generalization and reduce confusion between similar diseasesondifferentcrops.
4.1 Dataset
To ensure robust generalization, a comprehensive datasetwasaggregatedfrommultiplepublicsources:
• PlantVillage [23]: A foundational dataset with lab conditionimages.
•RiceLeafDiseases[10]:In-fieldimagesofcommon ricediseases.
•CassavaDiseases[10]:Imagesofcassavadiseases fromreal-worldconditions.
• PlantDoc [17]: Images of various plant diseases capturedwithmobiledevices.Thecombineddataset contains 53,303 images, covering44 unique cropdiseaseclassesacross12differentcroptypes.Images wereresizedto256×256pixels.
Table -1: DatasetComposition
4.2. Implementation Details
ThemodelwasimplementedusingTensorFlowand Keras.
• Hardware: NVIDIATeslaT4GPU(16GBVRAM).
• Optimizer: Adamwithaninitiallearningrateof10−3 and betas of (0.9,0.999). A ‘Reduce LR On Plateau‘ callback was used to decrease the learning rate if validationlossstagnated.
• Loss Function: Aweightedsumofcategoricalcross entropylossesforbothheads,withequalweightsof1.0.
• Regularization: Dropout(0.5inthesharedblock,0.3 intheheads)andL2weightdecay(10−5)wereusedto preventoverfitting.
• Batch Size: 32.
• Epochs: Trainedforamaximumof10epochs,withan earlystoppingcallbackmonitoringvalidationaccuracy withapatienceof8epochs.
• Augmentation: On- the- fly data augmentation includedrandomhorizontalandverticalflips,random rotation(±20°),zoom,andcontrastadjustment.

International Research Journal of Engineering and Technology (IRJET) e-ISSN: 2395-0056
Volume: 13 Issue: 01 | Jan 2026 www.irjet.net p-ISSN: 2395-0072
5.1. Comparison with State-of-the-Art
MS-HAN establishes new SOTA performance on the aggregatedmulti-cropdataset(seeTableII).
94.78%combinedaccuracy,outperformingEfficientNetB4 by1.66%.
•96.2%cropclassificationaccuracy.
•95.4%diseaseclassificationaccuracy.
•4.8Mparameters,makingitsignificantlymorelightweight thanmodelslikeResNet-50(25.6M)andViT-B/16(85.8M).
• Our model is over 18 times lighter than ViT-B/16 while achievinghigheraccuracy,highlightingitsefficiency.
Table -2: Performance Comparison with State-Of-TheArt models
Model
(%) Parameters (M)
5.2. Ablation Study
Tovalidatethecontributionofeachproposedcomponent, weconductedanablationstudy(seeTableIII).Westarted withaResNet-18asabaselineandincrementallyaddedour proposedmodules.
The hierarchical attention mechanism further improvedaccuracyby+2.3%byfocusingthemodelon salientregions.
The multi-scale module provided a significant boost of+3.2%overthebaseline,confirmingitseffectiveness incapturingvariedfeatures.
Decomposing the problem with the multi-head architectureyieldedthelargestsingleimprovementof +3.5%, validating our hypothesis that separating the tasksaidsgeneralization.
The full MS-HAN model achieves a +10.1% total ac curacy improvement over the baseline while having lessthanhalftheparameters.
The multi-head architecture demonstrated significant advantagesoveratraditionalsingle-headapproach(which combinescropanddiseaseintooneclass,e.g.,’TomatoLate Blight)
Improved Generalization: Disease classification accuracy improved by 3.5% for diseases that appear acrossmultiplecrops(e.g.,PowderyMildew).
Reduced Confusion: The confusion matrix showed that misclassifications between similar diseases in different crops (e.g., Apple Scab vs. Grape Black Rot) decreasedby42%.
Faster Convergence: Themodelwithseparateheads con verged approximately 30% faster in terms of epochscomparedtoasingle-headvariant,likelydueto thedecomposedandsimplifiedlearningobjectives.
This paper introduced MS-HAN, a novel deep learning architecture for generalized plant disease detection. By integrating an Inception-style multi-scale module, a hierarchical attention mechanism, and a multi-head classificationframework,MS-HANeffectivelyaddressesthe critical challenges of scale variance and cross-crop generalization.Ourmodelachievesstate-of-the-artaccuracy (94.78%) on a large, diverse dataset while maintaining a lightweightdesign(4.8Mparameters)suitableforrealworld deployment on edge devices. The results confirm that a domain-specific architecture that explicitly models the hierarchical nature of the problem (crop, then disease) significantlyoutperformsgeneric,single-taskmodels.Future workwillfocusonintegratingenvironmentalmetadatainto themodel,expandingthedatasetwithmorecropvarieties, andexploringknowledgedistillationtechniquesforfurther modelcompression
This research was supported by the Department of ComputerEngineering,ShriG.S.InstituteofTechnologyand Science,Indore.TheauthorthanksProf.SurendraGuptafor

International Research Journal of Engineering and Technology (IRJET) e-ISSN: 2395-0056
Volume: 13 Issue: 01 | Jan 2026 www.irjet.net p-ISSN: 2395-0072
guidance, and Kaggle for providing the computational resourcesnecessaryfortraining.
[1]S.P.Mohanty,D.P.Hughes,andM.Salath´e,”Usingdeep learning for image-based plant disease detection,” Front. PlantSci.,vol.7,p.1419,2016.
[2] J. Pandian et al., ”Identification of plant leaf diseases using a 9 layer deep CNN,” IEEE Trans. Agric., vol. 45, pp. 112–120,2019.
[3] S. Woo et al., ”CBAM: Convolutional block attention module,”inECCV,2018,pp.3–19.
[4]J.Hu,L.Shen,S.Albanie,etal.,”Squeeze-and-excitation networks,”inCVPR,2018,pp.7132–7141.
[5]S.Zhangetal.,”Multi-scalefeaturefusionCNNforplant diseaserecognition,”PlantPhenomics,vol.2021,pp.1–10, 2021.
[6] K. He et al., ”Deep residual learning for image recognition,”inCVPR,2016,pp.770–778.
[7]M.TanandQ.Le,”EfficientNet:Rethinkingmodelscaling forCNNs,”inICML,2019,pp.6105–6114.
[8]A.Dosovitskiy etal.,”An imageis worth 16x16 words: Trans formers for image recognition,” arXiv:2010.11929, 2020.
[9] Z. Liu et al., ”Swin transformer: Hierarchical vision transformer using shifted windows,” in ICCV, 2021, pp. 10012–10022.
[10]H.Iranga,”Leafdiseasedataset:IntegratingPlantVillage, rice,andcassavaimages,”Kaggle,2021.
[11] FAO, ”The impact of plant diseases on food security,” FAO,2021.
[12]K.P.Ferentinos,”Deeplearningmodelsforplantdisease detection,”Comput.Electron.Agric.,vol.145,pp.311–318, 2018.
[13]A.KamilarisandF.X.Prenafeta-Boldu,”Deeplearningin agriculture:Asurvey,”Comput.Electron.Agric.,vol.147,pp. 70–90,2018.
[14]L.Wang,S.Deng,andY.Sun,”Areviewofdeeplearning inplantdiseasedetection,”Comput.Electron.Agric.,vol.178, p.105760,2020.
[15] U. Nawaz et al., ”AI in Agriculture: A Survey of Deep Learning Techniques for Crops,” arXiv:2507.22101, 2025.
[16]H.Brahimi,K.Bettahar,M.Boulidam,”Deeplearningfor
plant diseases: Detection and saliency map visualisation,” Humanoids,2018.
[17] S. Singh et al., ”PlantDoc: A Dataset for Visual Plant DiseaseDetection,”arXiv:2001.08501,2020.
[18] C. Szegedy et al., ”Going deeper with convolutions,” CVPR,pp.1–9,2015.
[19]Z.Chen,L.Li,etal.,”Pyramidfeaturefusionforriceleaf disease detection,” Comput. Electron. Agric., vol. 193, p. 106656,2022.
[20]H.Chengetal.,”Identificationofappleleafdiseasevia novelattentionmechanism,”FrontiersinPlantScience,vol. 14,p.1274231,2023.
[21]A.Pandey,S.Singh,A.Pathak,”Arobustdeepattention dense convolutional neural network for plant leaf disease classification,”ExpertSystemswithApplications,vol.205,p. 117676,2022.
[22] E. Zhang, N. Zhang, F. Li, C. Lv, ”A lightweight dualattention network for tomato leaf disease identification,” FrontiersinPlantScience,vol.15,p.1420584,2024.
[23]D.P.Hughes,M.Salath´e,”Anopenaccessrepositoryof imagesonplanthealthtoenablethedevelopmentofmobile diseasediagnostics,”arXivpreprintarXiv:1511.08060,2015.
[24]R.Bugubaeva,etal.,”DeepLabV3+forsegmentationof agricul tural images,” Computers and Electronics in Agriculture,vol.205,p.107695,2023.
[25] E. Shelhamer, J. Long, T. Darrell, ”Fully convolutional networksforsemanticsegmentation,”CVPR,pp.3431–3440, 2017

oto
AishwaryaMalviya, Department of Computer Engineering
Shri G.S. Institute of Technology and Science Indore- 452001, MadhyaPradesh,India
Email:aishmaliviya@gmail.com