Team Mutographs

The team that transformed understanding of how cancer starts.
Funded 2017-2024 by Challenge
Unusual mutation patterns

Dare to think differently


The team that transformed understanding of how cancer starts.
Funded 2017-2024 by Challenge
Unusual mutation patterns

Dare to think differently

Founded by

Cancer Grand Challenges is a global initiative that is building a pioneering, interdisciplinary community to take on and solve cancer’s most complex problems.
Co-founded by the two largest funders of cancer research in the world, Cancer Research UK and the National Cancer Institute in the US, Cancer Grand Challenges aims to accelerate high-impact research and translate discoveries for public and patient benefit by transforming how team science is conducted.
Through Cancer Research UK, Cancer Grand Challenges is building a network of like-minded partners and individual donors around the world, all of whom share our aspiration to create change. Our work wouldn’t be possible without their collective support.
In 2017, funded by Cancer Research UK, team Mutographs set out to tackle the unusual mutations patterns challenge, to discover how unusual patterns of mutation are induced by different cancer-causing events. By studying the tell-tale fingerprints etched into the DNA of both healthy and cancerous cells, the team aimed to identify unknown causes of the disease and shed light on geographical differences in cancer incidence.
When the challenge was set, the prevailing thinking in the field was that each disease risk factor or carcinogen would leave behind a distinctive pattern of mutations, known as a ‘mutational signature’. For example, cancers triggered by exposure to UV radiation will exhibit distinct signatures compared to those linked with tobacco use. Team Mutographs hypothesised that working backward from these mutational signatures could identify new, potentially preventable causes of cancer.

“The wonderful thing about Cancer Grand Challenges is its mindset. It encourages you to think big, combine different fields of expertise and not let operational obstacles get in the way of addressing important questions.”
Sir Mike Stratton
Mutographs
Team Lead, Wellcome Sanger Institute


The team that transformed understanding of how cancer starts. Mutographs brought together world-leading epidemiologists with computational biologists, geneticists and clinicians from the UK, US and France and connected them with cancer centres across the globe.
The team was led by Sir Mike Stratton, then Director of the Wellcome Sanger Institute in Cambridge, UK.

Map Key
United States
United Kingdom France
Stephen Elledge,

SPECIFICANCER Team Lead and Gregor Mendel Professor of Genetics, Harvard Medical School and Brigham and Women’s Hospital.
Sir Mike Stratton
Mutographs Team Lead, Wellcome Sanger Institute




















“Mutographs’
ambition was on an epic and global scale, involving the collection and genomic analysis of samples from thousands of cancer patients worldwide.”
Gemma Balmer-Kemp
Head of Research, Cancer Grand Challenges
Tissue samples collected
From:
> 40 medical centres
> 30 countries 5 continents
~15,000 >100,000
Number of genomes sequenced
~1,000,000,000,000,000
Bytes of data
By utilising its novel ‘mutational epidemiology’ approach at an unprecedented scale, the team set out to comprehensively catalogue mutational signatures present in the genomes of both cancerous and normal cells, and thereby uncover the underlying mechanisms behind cancer development.
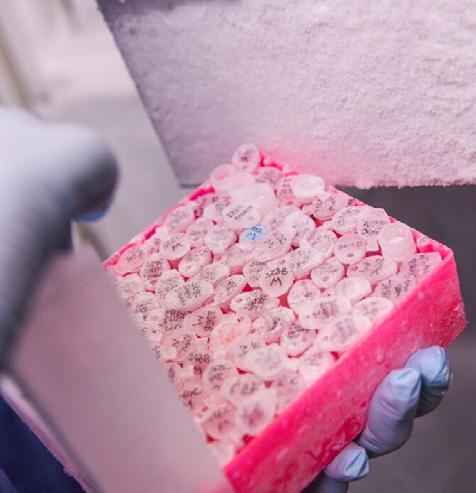
Thanks to a unique and pioneering strategic partnership between the Wellcome Sanger Institute and the International Agency for Research in Cancer (WHOIARC), enabled by Cancer Grand Challenges, the team gathered samples from 7,800 patients across eight cancer types – including pancreatic, kidney, oesophageal, bowel, head and neck, bladder and gallbladder. People were recruited from over 40 medical centres in more than 30 countries, spanning five continents. Critically, for each cancer type, patients were included from regions with high or low incidence rates. This extensive biorepository containing over 100,000 tissue samples along with detailed demographic, lifestyle, environmental, clinical data, and whole genome sequences, holds immense promise for future discoveries and will remain a valuable resource for the scientific community for years to come.
By studying this vast data, the team has uncovered several exposures linked to cancer incidence in multiple cancer types, which collectively affect potentially tens of millions of people. The team are working to determine the source of these exposures.
But through unexpected findings Mutographs has also made seminal discoveries that have transformed scientific understanding about the role of mutations in carcinogenesis. When the challenge was set, the predominant thinking in the field was that most carcinogens caused cancer by directly altering cellular DNA. However, by exposing mouse models to suspected carcinogens, the team observed that many carcinogens do not actually cause an increase in mutations, and presumably act via other means. In oesophageal squamous cell carcinoma the team also found no geographical variation in mutational signatures despite dramatic variance in global incidence rates. In addition, the team’s research has added to the body of work revealing that many healthy tissues harbour complex and diverse mutational landscapes. Together, these findings have led to a major shift in how the scientific community thinks about how carcinogens cause cancer, reigniting interest in the promotional hypothesis of carcinogenesis, by uncovering an important role of non-mutagenic mechanisms. The team has simultaneously emphasised the critical role of mutations, highlighting they are essential, but in some cases not sufficient to drive cancer.
Characterizing Mutational Signatures in Human Cancer Cell Lines Reveals Episodic APOBEC Mutagenesis CELL
Somatic
Tobacco smoking and somatic mutations in human bronchial epithelium NATURE
Mutographs awarded $25m in funding
When the team sequenced bronchial epithelial cells, it revealed that while smokers carry markedly elevated mutational burdens, a subset of cells retains normal mutation levels. Strikingly, this subset significantly expanded in ex-smokers. This suggests that cells spared from tobacco-induced mutagenesis actively repopulate the bronchial epithelium after individuals quit smoking. These findings provide a new perspective on lung regeneration as well as identifying variation in levels of mutagenesis due to tobacco smoke.
Uncovering the bigger picture
Timeline Key:
Understanding APOBEC mutagenesis
Reviving the promotional hypothesis
The repertoire of mutational signatures in human cancer NATURE
NORMAL PHENOTYPES CHALLENGE SET
The mutational signature profile of known and suspected human carcinogens in mice

Reliable detection of somatic mutations in solid tissues by
microdissection and low-input DNA sequencing
Inspired by Mutographs work showing that normal tissues can be riddled with oncogenic mutations yet remain phenotypically normal, Cancer Grand Challenges set the normal phenotypes challenge, to understand how cells and tissues maintain normality and how they then transition to becoming a tumour. The PROMINENT team was subsequently funded to address the challenge.
To enable reliable detection of somatic mutations in solid tissues, which are typically heterogenous and often polyclonal, team members at the Sanger developed laser-capture microdissection and low-input DNA sequencing. This allows whole genome sequencing of small populations of cells whilst preserving tissue architecture, giving phenotypic and spatial information.
Increased somatic mutation burdens in normal human cells due to defective DNA polymerases
NATURE GENETICS
It was thought the ageing phenotype is driven by the accumulation of somatic mutations in cells. By analysing tissues from individuals with inherited cancer predisposition syndromes, the team showed that although these individuals had elevated mutation rates in most normal cells, causing cancer susceptibility, they didn’t show other obvious ageing phenotypes. This indicated that the body could tolerate higher mutation rates and suggested that the somatic mutational theory of ageing was more complex than just the accumulation of mutations driving the ageing phenotype, challenging another longstanding dogma of the field.

Convergent somatic mutations in metabolism genes in chronic liver disease
NATURE

In liver, the team detailed the mutational signatures of normal hepatocytes, and when sequencing individuals with liver diseases observed mutations in key metabolism regulatory genes. These were acquired in multiple clones independently, suggesting convergent evolution, with a new class of “driver” mutation providing a protective mechanism against lipotoxicity associated with liver disease. This work highlights the team’s approach can be used to discover biological pathways that mitigate the effects of systemic diseases, and be used to identify targets that could be exploited therapeutically.

First author and Mutographs future leader Stanley W.K. Ng is now an Assistant Professor at the University of California, Irvine, US.

Mutational signatures in esophageal squamous cell carcinoma from eight countries with varying incidence
NATURE
Mapping clustered mutations in cancer reveals APOBEC3 mutagenesis of ecDNA
NATURE
Mechanisms of APOBEC3 mutagenesis in human cancer cells
NATURE
Topography of mutational signatures in human cancer CELL REPORTS
QUOTIENT THERAPEUTICS
Multiple seminal papers from the team come together to change the way we think about carcinogenesis. In the classical view, carcinogens cause mutations, and these mutations cause cancer. But the team demonstrated that normal tissues have diverse mutational profiles, already carrying oncogenic mutations. Mutographs went on to show in mouse models that 17/20 suspected carcinogens did not cause mutations, despite promoting tumour formation. The team’s findings in oesophageal squamous cell carcinoma then found no mutational signatures that could account for the extreme variance in global incidence rates. Together, Mutographs work has highlighted the importance of nonmutagenic mechanisms of cancer promotion.
The team uncovered that APOBEC mutagenesis occurs in episodic bursts, established its causal role in generating mutational signatures SBS2 and SBS13, and identified APOBEC3A as the main family member driving these processes. Mutographs also revealed a role of APOBEC3 activity in extrachromosomal DNA mutagenesis, driving its evolution.

First author and Mutographs future leader Mia Petljak is now an Assistant Professor at the NYU Grossman School of Medicine, US.

First author and Mutographs future leader Burçak Otlu is now an Assistant Professor at the Middle East Technical University, Turkey.

Quotient Therapeutics launched
Mike Stratton, Inigo Martincorena and Peter Campbell are academic co-founders’.
The Mutographs biorepository: A unique genomic resource to study cancer around the
Mutographs revealed a rich landscape of exposures accounting for geographical variance in clear cell renal cell carcinoma, demonstrating the power of its mutational epidemiology approach. The team identified a new widespread signature, SBS40b, which potentially explains the variation in incidence across the world, with mutation loads correlating with incidence rates. Mutographs also found a new signature unique to Japan, SBS12, and that exposure to aristolochic acids was much more widespread than previously thought in the Balkan region.
Implementing Mutational Epidemiology on a Global Scale: Lessons from Mutographs CANCER DISCOVERY
The complexity of tobacco smoke-induced mutagenesis in head and neck cancer NATURE GENTICS Geographic

By studying 265 head and neck cancer samples from patients across eight countries, the team identified six mutational signatures associated with tobacco smoke and at least three different mutational processes underlying them. Improved sampling resolution showed variation of these processes across anatomical sites. The team went on to show an association between smoking and alcohol related signatures, indicating that these risk factors can combine to cause mutations.

Geographic and age variations in mutational processes in

By analysing 981 colorectal cancer genomes from patients across 11 countries, Mutographs identified mutational signatures derived from the bacterial genotoxin colibactin, produced by E. coli harbouring the pks operon. The signatures, SBS88 and ID18, correlated with cancer incidence across the globe and were also enriched in early onset colorectal cancer. Intriguingly, the signatures were imprinted early in colorectal cancer development and linked to APC driver mutations. The team also found mutational signatures of unknown origin, which were more prevalent in South America. One of them, SBS89, shares many of the characteristics of the colibactin derived signature SBS88. This potentially indicates the existence of multiple kinds of mutagenic microbes found in different locations that could play a role in colorectal cancer and thereby explain some of the differences with respect to geography. Mutographs work has thus far shown a correlation between early-life exposure to mutagenic bacteria and early onset colorectal cancer, which is rising at an alarming rise in those under the age of 50. However, more work is needed to demonstrate a causal link.

First author and Mutographs future leader Marcos
Gay is now a Junior Group Leader at the Spanish National Cancer Research Center (CNIO), Spain.
NATURE GENETICS JULY 2025
The long-term effects of chemotherapy on normal blood cells
The team carried out the first systematic analysis of the mutational impact of chemotherapy on normal tissues. By sequencing blood cell genomes from 23 patients aged 3–80 treated with a range of chemotherapies, Mutographs identified different chemotherapies induce different mutational signatures, as well as four new mutational signatures. Certain chemotherapies, including some platinum agents, induced mutation burdens in young patients comparable to those of elderly individuals, and caused premature changes in hematopoietic stem cell populations similar to natural ageing. The team hopes its work will guide treatment selection to deliver effective chemotherapies with far fewer damaging side effects for patients.

First author and Mutographs future leader Emily Mitchell is now a Consultant in Haematology at the Cambridge University Hospitals NHS Foundation Trust, UK.
In addressing the unusual mutations pattern challenge, the Mutographs team uncovered new mutational signatures, some of which, have the potential to explain the global variance in cancer incidence. When the team set out to address the challenge, approximately 30 mutational signatures arising from single base substitutions (SBS) had been identified. As of March 2025, there were at least 86 single-base pair and dinucleotide mutational signatures catalogued in the COSMIC database, many of which were identified by the team. Inspired by Mutographs work, Cancer Grand Challenges set the mechanisms driving mutational signatures challenge, to understand the molecular processes or exposures that generate unexplained mutational signatures, and ultimately to move the field towards actionable insights for cancer prevention.

...with work on breast, pancreas and gall-bladder still to come


‘The team’s findings showing that many carcinogens don’t actually induce DNA mutations totally changed our understanding of how cancer starts in the first place. That is a hugely important finding, because it completely opens up a whole new opportunity in the field of cancer prevention. The team has changed our fundamental view on cancer.’
Charles Swanton Chair of the Cancer Grand Challenges Scientific Committee
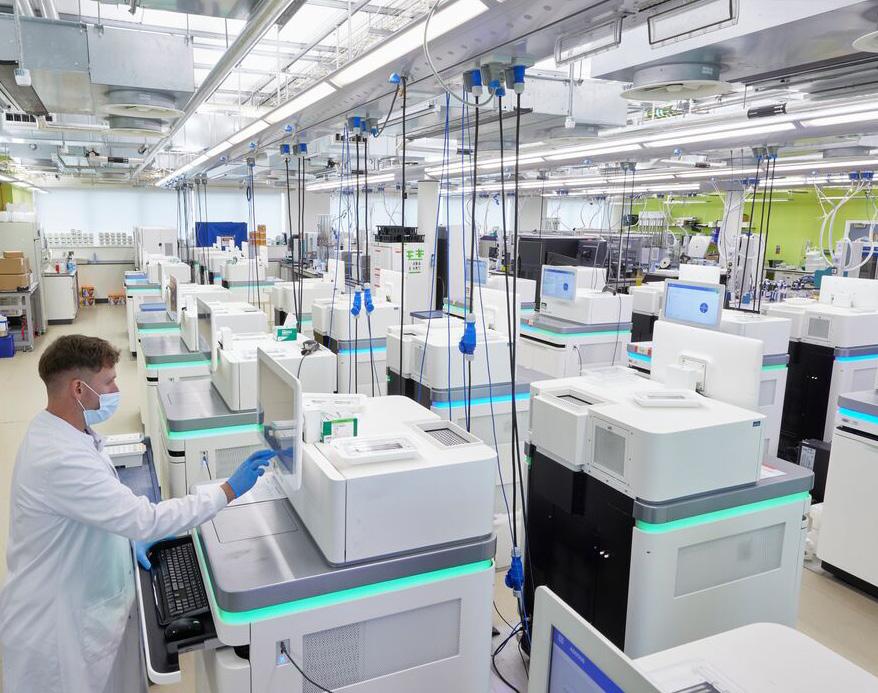
To enable its groundbreaking discoveries, the team developed a suite of AI-powered tools to identify and analyse mutational signatures. The team has made all of these tools widely available to the scientific community. This includes SigProfiler and all of its variants (Extractor, Topography, Clusters, Plotting, Matrix Generator, Simulator, Assignment), as well as the Mutational Signature Attribution (MSA) tool. These tools have already been extensively adopted, adding to the team’s scientific legacy that will drive further discoveries. The team has also enhanced the provision of the widely used Catalogue of Somatic Mutations in Cancer (COSMIC) web portal, adding more than forty mutational signatures and other new features.
Mutographs trainees have become members of a new generation of cancer researchers where ideas are constantly exchanged across time zones and geographical regions in order to solve the most complex and pertinent questions in cancer biology. This continued interaction has led to previously unseen global collaborations, forging long-lasting scientific relationships. Cancer Grand Challenges funding has enabled the Mutographs team to train over 15 PhD students and 25 postdoctoral researchers, with eight going on to establish their own independent research groups so far.
Patient advocates are embedded into Cancer Grand Challenges teams and are important members of the teams we fund to address cancer’s toughest challenges. The Mutographs team included two patient advocates: Mimi McCord and Maggie Blanks. They initially focused on gaining an understanding of the cultural, socioeconomic and societal contexts of cancer care and research in some of the countries where the team was collecting samples, to inform on how to best communicate any relevant healthcare messages. Mimi and Maggie then worked alongside the researchers to create lay language documents, review data collection documentation and brought about the inclusion of the patients’ voice in ethical review discussions of partnering institutions. The initiative is learning from the inaugural teams’ experiences, both the advocates and researchers, to ensure the patient voice and experience is at the heart of our work.
$25m 8 15 8 38 25
Number of
Number of postdoctoral researchers trained

Team Mutographs has pioneered the field of mutational epidemiology, paving the way for potential new approaches to cancer prevention.
The team’s groundbreaking discoveries range from identifying unknown mutational signatures and uncovering multiple widespread mutagenic exposures of known and unknown causes, to revealing the complex mutation patterns in healthy tissues, challenging the somatic theory of ageing, as well as adding to the body of evidence for the promotional theory of carcinogenesis. The team went on to uncover clues as to why global incidence rates vary for some cancer types, the potential role of the microbiome in early-onset bowel cancer, the interplay of alcohol and tobacco in cancer risk and the mutational effects of chemotherapies on normal tissues.
Mutographs’ findings have fed into the work of multiple other teams and directly led to Cancer Grand Challenges setting both the normal phenotypes challenge in 2020 and the mechanisms driving mutational signatures challenge in 2025. In 2022, the PROMINENT team was funded to investigate how cells maintain a normal phenotype despite harbouring cancer causing mutations and what causes them to become cancerous.
Researchers from Mutographs continue to work towards determining the source of the unknown exposures causing the identified mutational signatures and further expanding their mutational epidemiology approach, toward a complete global catalogue of mutational signatures
The team’s findings have changed the way we think about cancer, demonstrating the power of global team science. Together with the tools and techniques developed, the vast biorepository created and the future leaders trained they form a legacy that continues to change thinking around how cancer starts and will accelerate routes to cancer prevention.
Funded institutes:
International Agency for Research on Cancer King’s College London
UCSF Helen Diller Family Comprehensive Cancer Center
University of California San Diego
Wellcome Sanger Institute
